Stanで、負の二項分布へのあてはめと予測区間の描画 [統計]
Rのglm.nb()では予測区間を求めるのがけっこう面倒そうなので、Stanでやってみたメモです。
Rコード
## Data
N <- 200
NB <- 10
m1 <- 0
m2 <- 0
sd1 <- 0.2
sd2 <- 0.3
sd.B <- 0.8
intercept <- 1
coef.X1 <- 1
set.seed(123)
X1 <- rnorm(N, 0, sd1)
X2 <- rnorm(N, 0, sd2)
B <- rep(rnorm(NB, 0, sd.B), each = N / NB)
Y <- rpois(N, exp(intercept + coef.X1 * X1 + B))
library(rstan)
rstan_options(auto_write = TRUE)
options(mc.cores = parallel::detectCores())
newx <- seq(-0.8, 0.8, length = 100)
stan.data <- list(N = N,
X1 = X1,
X2 = X2,
Y = Y,
K = length(newx),
New_X1 = newx)
fit.stan <- stan("overdispersion_glm.stan",
data = stan.data,
iter = 2000, warmup = 1000, thin = 1)
print(fit.stan)
## Expectations
alpha <- get_posterior_mean(fit.stan,
pars = "alpha")[, "mean-all chains"]
beta1 <- get_posterior_mean(fit.stan,
pars = "beta")["beta[1]", "mean-all chains"]
exp.y <- exp(alpha + beta1 * newx)
## Prediction intervals
yy <- extract(fit.stan, pars = "y")
low10 <- sapply(seq_along(newx),
function(i) quantile(yy[[1]][, i], probs = 0.10))
low25 <- sapply(seq_along(newx),
function(i) quantile(yy[[1]][, i], probs = 0.25))
up75 <- sapply(seq_along(newx),
function(i) quantile(yy[[1]][, i], probs = 0.75))
up90 <- sapply(seq_along(newx),
function(i) quantile(yy[[1]][, i], probs = 0.90))
## plot
png()
plot(Y ~ X1, las = 1)
lines(newx, exp.y, lty = 1)
lines(newx, low25, lty = 2)
lines(newx, up75, lty = 2)
lines(newx, low10, lty = 3)
lines(newx, up90, lty = 3)
dev.off()
Stanコード
data {
int<lower=0> N;
vector[N] X1;
vector[N] X2;
int<lower=0> Y[N];
int<lower=0> K;
vector[K] New_X1;
}
parameters {
real alpha;
vector[2] beta;
real<lower=0> phi;
}
model {
vector[N] eta = alpha + beta[1] * X1 + beta[2] * X2;
phi ~ cauchy(0, 5);
Y ~ neg_binomial_2_log(eta, phi);
}
generated quantities {
vector[K] y;
for (i in 1:K) {
real mu = exp(alpha + beta[1] * New_X1[i]);
y[i] = neg_binomial_2_rng(mu, phi);
}
}
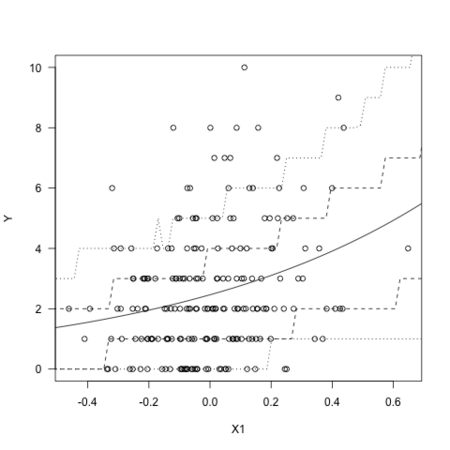
結果のグラフです。X2は0として、X1の変化に対するYの期待値と50%・80%予測区間です。





コメント 0