折れ線へのあてはめ [統計]
下のグラフのような折れ線状のデータへのあてはめをおこなうモデルをかんがえる。変化点の位置も不明とする。
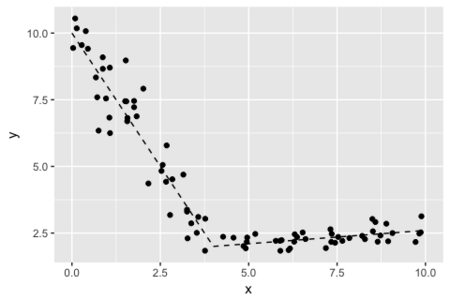
データを生成するRコード。
## Data
library(ggplot2)
set.seed(12)
n <- 40
cp <- 4
y0 <- 10
s1 <- -2
s2 <- 0.1
x1 <- runif(n, 0, cp)
x2 <- runif(n, cp, 10)
sd1 <- 1
sd2 <- 0.3
y1 <- rnorm(n, y0 + s1 * x1, sd1)
y2 <- rnorm(n, (y0 + s1 * cp) + s2 * (x2 - cp), sd2)
x <- c(x1, x2)
y <- c(y1, y2)
ggplot(data.frame(x = x, y = y)) +
geom_point(aes(x = x, y = y)) +
geom_segment(data = data.frame(x = 0, y = y0, xend = cp, yend = y0 + s1 * cp),
aes(x = x, y = y, xend = xend, yend = yend),
linetype = 2) +
geom_segment(data = data.frame(x = cp, xend = 10,
y = y0 + s1 * cp,
yend = y0 + s1 * cp + s2 * (10 - cp)),
aes(x = x, y = y, xend = xend, yend = yend),
linetype = 2)
まずStanのモデル。
data {
int N;
vector[N] X;
vector[N] Y;
}
transformed data {
real max_X = max(X);
real min_X = min(Y);
}
parameters {
vector[3] beta;
vector<lower=0>[2] sigma;
real<lower=min_X,upper=max_X> cp_x; // Change point
}
transformed parameters {
real cp_y = beta[1] + beta[2] * cp_x; // Y at the change point
}
model {
cp_x ~ uniform(min_X, max_X);
for (i in 1:N)
if (X[i] < cp_x)
Y[i] ~ normal(beta[1] + beta[2] * X[i], sigma[1]);
else
Y[i] ~ normal(cp_y + beta[3] * (X[i] - cp_x), sigma[2]);
}
JAGSのモデル。
var
N,
X[N],
Y[N],
cp_x,
max_x,
min_x,
beta[3],
sigma[2],
tau[2],
mu[N],
t[N];
data {
max_x <- max(X)
min_x <- min(X)
}
model {
for (i in 1:N) {
mu[i] <- (1 - step(X[i] - cp_x)) * (beta[1] + beta[2] * X[i]) +
step(X[i] - cp_x) * (beta[1] + beta[2] * cp_x + beta[3] * (X[i] - cp_x))
t[i] <- (1 - step(X[i] - cp_x)) * tau[1] +
step(X[i] - cp_x) * tau[2]
Y[i] ~ dnorm(mu[i], t[i])
}
cp_x ~ dunif(min_x, max_x)
for (i in 1:3) {
beta[i] ~ dnorm(0, 1.0e-2)
}
for (i in 1:2) {
sigma[i] ~ dunif(0, 1.0e+4)
tau[i] <- 1 / (sigma[i] * sigma[i])
}
}
StanとJAGSを実行するRコード。
## Stan
library(rstan)
rstan_options(auto_write = TRUE)
options(mc.cores = parallel::detectCores())
data <- list(N = n * 2,
X = x,
Y = y)
stan.out <- stan("cp.stan", data = data,
pars = c("cp_x", "beta", "sigma"),
control = list(max_treedepth = 15))
print(stan.out)
## JAGS
library(rjags)
data <- list(N = n * 2,
X = x,
Y = y)
model <- jags.model("cp_bug.txt", data = data,
n.chains = 4, n.adapt = 1000)
update(model, n.iter = 1000)
jags.out <- coda.samples(model,
variable.names = c("cp_x", "beta", "sigma"),
n.iter = 2000)
gelman.diag(jags.out)
summary(jags.out)
結果です。
> print(stan.out)
Inference for Stan model: cp.
4 chains, each with iter=2000; warmup=1000; thin=1;
post-warmup draws per chain=1000, total post-warmup draws=4000.
mean se_mean sd 2.5% 25% 50% 75% 97.5% n_eff Rhat
cp_x 4.00 0.00 0.14 3.79 3.90 3.99 4.09 4.30 1478 1
beta[1] 10.07 0.01 0.27 9.52 9.89 10.07 10.25 10.59 1679 1
beta[2] -2.01 0.00 0.12 -2.22 -2.09 -2.01 -1.93 -1.76 1316 1
beta[3] 0.09 0.00 0.03 0.04 0.07 0.09 0.11 0.14 2108 1
sigma[1] 0.94 0.00 0.11 0.75 0.86 0.92 1.00 1.18 3024 1
sigma[2] 0.27 0.00 0.03 0.22 0.25 0.27 0.29 0.35 2036 1
lp__ 15.72 0.05 1.80 11.26 14.76 16.07 17.06 18.16 1537 1
> gelman.diag(jags.out)
Potential scale reduction factors:
Point est. Upper C.I.
beta[1] 1.04 1.11
beta[2] 1.05 1.15
beta[3] 1.00 1.01
cp_x 1.05 1.13
sigma[1] 1.00 1.01
sigma[2] 1.00 1.00
Multivariate psrf
1.05
> summary(jags.out)
Iterations = 2001:4000
Thinning interval = 1
Number of chains = 4
Sample size per chain = 2000
1. Empirical mean and standard deviation for each variable,
plus standard error of the mean:
Mean SD Naive SE Time-series SE
beta[1] 10.06806 0.29514 0.0032997 0.0329236
beta[2] -2.00325 0.12333 0.0013788 0.0205014
beta[3] 0.09067 0.02623 0.0002933 0.0008305
cp_x 4.00965 0.13609 0.0015215 0.0148817
sigma[1] 0.93673 0.11151 0.0012467 0.0017457
sigma[2] 0.26778 0.03219 0.0003600 0.0004189
2. Quantiles for each variable:
2.5% 25% 50% 75% 97.5%
beta[1] 9.43796 9.8798 10.07672 10.2679 10.6282
beta[2] -2.24950 -2.0832 -2.00775 -1.9208 -1.7542
beta[3] 0.03913 0.0734 0.09059 0.1085 0.1417
cp_x 3.78724 3.9082 3.99859 4.1002 4.3023
sigma[1] 0.74820 0.8598 0.92435 1.0032 1.1862
sigma[2] 0.21443 0.2449 0.26461 0.2873 0.3385
2017-09-18 08:31
nice!(1)
コメント(0)





コメント 0